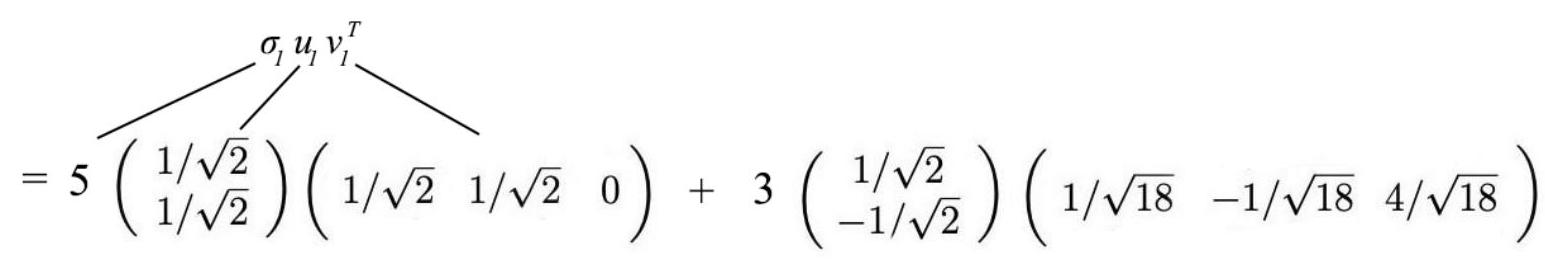| height | weight | |
|---|---|---|
| height | 4.86 | 6.67 |
| weight | 6.67 | 9.74 |
Ch 5 Lecture 3
Intuition about SVD
Thinking again about matrix diagonalization
The same logic, applied to a general matrix
Recap
A=U S V^{T}
where
\begin{array}{lccccc} & & & U & V & S \\ & & &\left(\begin{array}{ccc} \mid & & \mid \\ \mathbf{u}_{1} & \cdots & \mathbf{u}_{m} \\ \mid & & \mid \end{array}\right) &\left(\begin{array}{ccc} \mid & & \mid \\ \mathbf{v}_{1} & \cdots & \mathbf{v}_{n} \\ \mid & & \mid \end{array}\right) &\left(\begin{array}{cccccc} \sigma_1 & & & & & \\ & \sigma_2 & & & & \\ & & \ddots & & & \\ & & & \sigma_r & & \\ & & & & \ddots & \\ & & & & & 0 \end{array}\right)\\ \text{eigenvectors of} & & & A A^{T} & A^{T} A \\ \end{array}
Example
A=\left(\begin{array}{ccc} 3 & 2 & 2 \\ 2 & 3 & -2 \end{array}\right)
A A^{T}=\left(\begin{array}{cc} 17 & 8 \\ 8 & 17 \end{array}\right), \quad A^{T} A=\left(\begin{array}{ccc} 13 & 12 & 2 \\ 12 & 13 & -2 \\ 2 & -2 & 8 \end{array}\right)
\begin{array}{cc} A A^{T}=\left(\begin{array}{cc} 17 & 8 \\ 8 & 17 \end{array}\right) & A^{T} A=\left(\begin{array}{ccc} 13 & 12 & 2 \\ 12 & 13 & -2 \\ 2 & -2 & 8 \end{array}\right) \\ \begin{array}{c} \text { eigenvalues: } \lambda_{1}=25, \lambda_{2}=9 \\ \text { eigenvectors } \end{array} & \begin{array}{c} \text { eigenvalues: } \lambda_{1}=25, \lambda_{2}=9, \lambda_{3}=0 \\ \text { eigenvectors } \end{array} \\ u_{1}=\binom{1 / \sqrt{2}}{1 / \sqrt{2}} \quad u_{2}=\binom{1 / \sqrt{2}}{-1 / \sqrt{2}} & v_{1}=\left(\begin{array}{c} 1 / \sqrt{2} \\ 1 / \sqrt{2} \\ 0 \end{array}\right) \quad v_{2}=\left(\begin{array}{c} 1 / \sqrt{18} \\ -1 / \sqrt{18} \\ 4 / \sqrt{18} \end{array}\right) \quad v_{3}=\left(\begin{array}{c} 2 / 3 \\ -2 / 3 \\ -1 / 3 \end{array}\right) \end{array}
SVD decomposition of A: A=U S V^{T}=\left(\begin{array}{cc} 1 / \sqrt{2} & 1 / \sqrt{2} \\ 1 / \sqrt{2} & -1 / \sqrt{2} \end{array}\right)\left(\begin{array}{ccc} 5 & 0 & 0 \\ 0 & 3 & 0 \end{array}\right)\left(\begin{array}{rrr} 1 / \sqrt{2} & 1 / \sqrt{2} & 0 \\ 1 / \sqrt{18} & -1 / \sqrt{18} & 4 / \sqrt{18} \\ 2 / 3 & -2 / 3 & -1 / 3 \end{array}\right)
Reformulating SVD
SVD as a sum of rank-one matrices
A=\sigma_{l} u_{l} v_{l}^{T}+\ldots+\sigma_{r} u_{r} v_{r}^{T}
Back to our example
A=\left(\begin{array}{ccc} 3 & 2 & 2 \\ 2 & 3 & -2 \end{array}\right)
A=\left(\begin{array}{ccc}3 & 2 & 2 \\ 2 & 3 & -2\end{array}\right)
Uses of SVD
Moore–Penrose pseudoinverse
Suppose we want to solve a linear system
A x = b.
If A is invertible, the solution is
x = A^{-1} b.
But many matrices are not invertible:
- A may not be square,
- or it may have linearly dependent columns.
In this case, an exact solution may not exist. Instead, we look for the vector x that minimizes the least–squares error
|Ax-b|_2.
Among all least–squares solutions, we choose the one with smallest norm.
This motivates the Moore–Penrose pseudoinverse A^+, defined so that
x = A^+ b
is the minimum-norm least–squares solution.
Using the SVD
Take the singular value decomposition
A = U \Sigma V^T,
where
- U and V are orthogonal matrices,
- \Sigma is diagonal (possibly rectangular):
\Sigma = \begin{bmatrix} \sigma_1 &&&\\ & \ddots &&\\ && \sigma_r &&\\ &&& 0 \end{bmatrix}, \qquad \sigma_1 \ge \cdots \ge \sigma_r >0.
Normal equations
Least squares solutions satisfy the Normal equations
A^T A x = A^T b.
Substitute the SVD:
(V \Sigma^T U^T)(U \Sigma V^T)x = V \Sigma^T U^T b.
Since U^TU=I,
V \Sigma^T \Sigma V^T x = V \Sigma^T U^T b.
Multiply on the left by V^T:
\Sigma^T \Sigma V^T x = \Sigma^T U^T b.
Solve componentwise
\Sigma^T \Sigma V^T x = \Sigma^T U^T b.
We can write this componentwise as
\Sigma^T \Sigma \begin{bmatrix} \mathbf{v}_1^T \mathbf{x} \\ \mathbf{v}_2^T \mathbf{x} \\ \vdots \\ \mathbf{v}_n^T \mathbf{x} \end{bmatrix}= \Sigma^T \begin{bmatrix} \mathbf{u}_1^T \mathbf{b} \\ \mathbf{u}_2^T \mathbf{b} \\ \vdots \\ \mathbf{u}_n^T \mathbf{b} \end{bmatrix}.
Since both \Sigma^T and \Sigma^T\Sigma are diagonal, they just multiply each element by the corresponding singular value (or square):
\Sigma^T \Sigma V^T \mathbf{x} = \Sigma^T U^T \mathbf{b}
becomes
\begin{bmatrix} \sigma_1^2 \mathbf{v}_1^T \mathbf{x} \\ \sigma_2^2 \mathbf{v}_2^T \mathbf{x} \\ \vdots \\ \sigma_n^2 \mathbf{v}_n^T \mathbf{x} \\ \end{bmatrix} = \begin{bmatrix} \sigma_1\mathbf{u}_1^T \mathbf{b} \\ \sigma_2\mathbf{u}_2^T \mathbf{b} \\ \vdots \\ \sigma_n\mathbf{u}_n^T \mathbf{b} \end{bmatrix}.
In other words, we get a separate equation for each singular value \sigma_i:
\sigma_i^2 \mathbf{v}_i^T \mathbf{x} = \sigma_i (\mathbf{u}_i^T \mathbf{b}).
Case 1: \sigma_i \neq 0
\sigma_i^2 \mathbf{v}_i^T \mathbf{x} = \sigma_i (\mathbf{u}_i^T \mathbf{b}).
We can divide by \sigma_i^2:
\mathbf{v}_i^T \mathbf{x} = \frac{1}{\sigma_i} \mathbf{u}_i^T \mathbf{b}.
Case 2: \sigma_i = 0
\sigma_i^2 \mathbf{v}_i^T \mathbf{x} = \sigma_i (\mathbf{u}_i^T \mathbf{b}).
The equation becomes
0 = 0,
which imposes no constraint on y_i.
To obtain the minimum-norm solution, we choose \mathbf{x} to be orthogonal to \mathbf{v}_i, so that:
\mathbf{v}_i^T \mathbf{x} = 0.
Defining \Sigma^+
These two cases can be summarized by defining the pseudoinverse of \Sigma
\Sigma^+ = \begin{bmatrix} 1/\sigma_1 &&&\\ & \ddots &&\\ && 1/\sigma_r &\\ &&& 0 \end{bmatrix}.
That’s just replacing every non-zero singular value with its reciprocal.
Then the equation becomes
V^T \mathbf{x} = \Sigma^+ U^T \mathbf{b}.
The Moore–Penrose pseudoinverse
V^T \mathbf{x} = \Sigma^+ U^T \mathbf{b}.
Finally, we can multiply both sides on the left by V to get
\mathbf{x} = V \Sigma^+ U^T \mathbf{b}
The quantity V \Sigma^+ U^T is acting like an inverse here – it is called the Moore–Penrose pseudoinverse of A.
For any vector b,
\mathbf{x} = A^+ \mathbf{b}
is the minimum-norm solution to the least-squares problem.
Interpretation
\mathbf{x} = V \Sigma^+ U^T \mathbf{b}
The SVD reveals exactly what the pseudoinverse does:
- Each singular direction acts independently.
- Directions with \sigma_i>0 can be inverted.
- Directions with \sigma_i=0 contain information lost by A and cannot be recovered.
The pseudoinverse therefore
undoes the action of A wherever inversion is possible but ignores directions where information has been destroyed.
:::
Example
Matrices as data
Example: Height and weight
A^T=\left[\begin{array}{rrrrrrrrrrrr}2.9 & -1.5 & 0.1 & -1.0 & 2.1 & -4.0 & -2.0 & 2.2 & 0.2 & 2.0 & 1.5 & -2.5 \\ 4.0 & -0.9 & 0.0 & -1.0 & 3.0 & -5.0 & -3.5 & 2.6 & 1.0 & 3.5 & 1.0 & -4.7\end{array}\right]
Why does A^T A appear? (Data interpretation)
Up to now, matrices like A^T A have appeared naturally during the construction of the SVD.
But suppose now that A is a data matrix.
What does A^T A actually represent?
Step 1 — Interpret A as data
Think of
- rows = observations (people)
- columns = variables (height, weight, etc.)
Each row is one data point in \mathbb{R}^p.
Step 2 — Center the data
Subtract the mean of each column so every variable has mean 0.
We now assume:
A \text{ is zero-centered.}
This assumption is the key bridge between linear algebra and statistics.
Step 3 — Examine one entry of A^T A
The (i,j) entry is
(A^T A)*{ij} = \sum*{k=1}^{n} A_{k i}A_{k j}.
This computation:
- multiplies variable i and variable j together,
- then adds across all observations.
That is exactly how we measure co-variation between variables.
Step 4 — The covariance matrix appears
For zero-centered data,
\Sigma = \frac{1}{n-1}A^T A
is the sample covariance matrix.
So the matrix that appeared in the SVD derivation is not mysterious at all:
A^T A measures how the variables vary together.
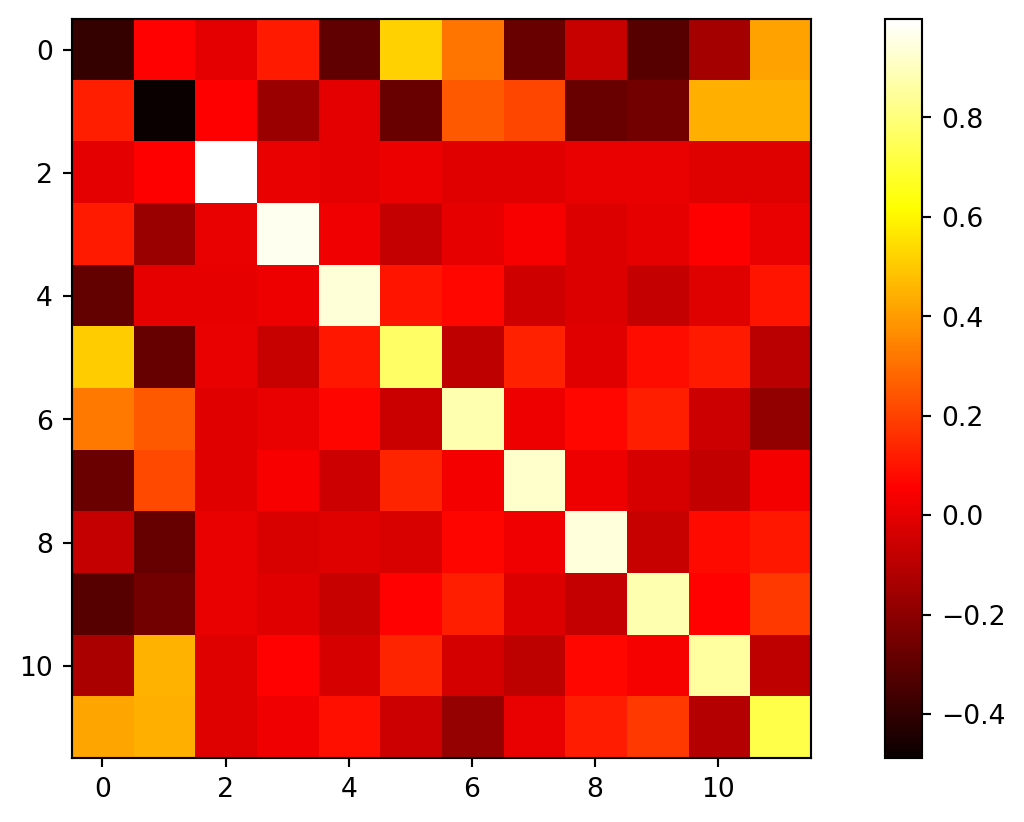
What do off-diagonal entries mean?
Look at a covariance matrix
\Sigma= \begin{bmatrix} \sigma_1^2 & c_{12} & \cdots c_{12} & \sigma_2^2 & \cdots \vdots & \vdots & \ddots \end{bmatrix}.
- The diagonal entries record how much each variable varies on its own.
- The off-diagonal entries measure how two variables move together.
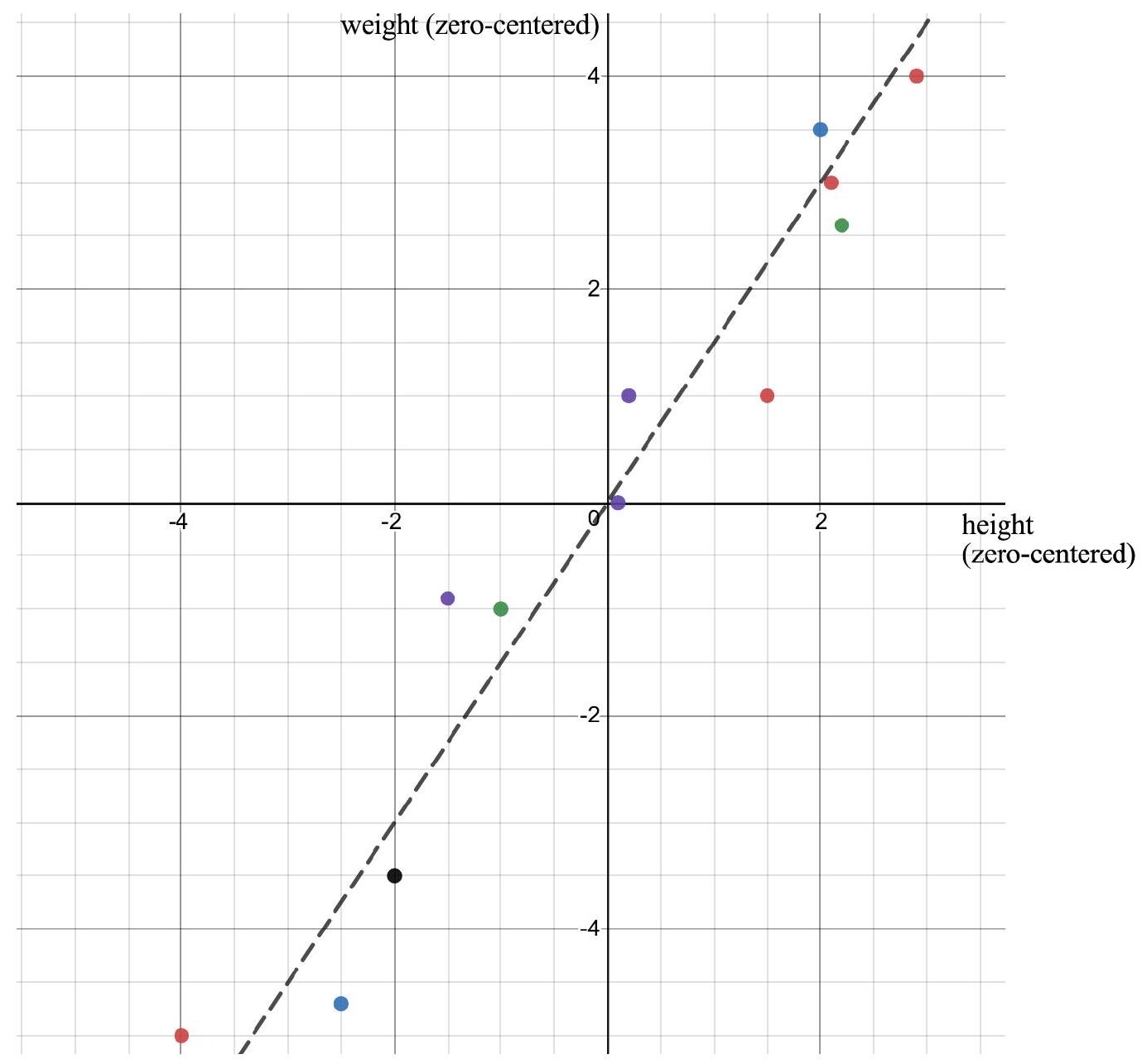
Heights and weights example
Let’s look at the (zero-centered) heights/weights dataset we’ve been using.
The off-diagonal entry \Sigma_{12}= 6.67 is the covariance between height and weight.
The positive value of the off-diagonal entry means that (in this dataset) when height increases, weight tends to increase too.
Knowin the height of an individual lets us make a guess about their weight.
But what if we rotated the axes?
A tilted data cloud suggests we are using a coordinate system that is not aligned with the natural shape of the data.
Let’s rotate to new axes:
In the rotated coordinate system, moving along one new axis does not force movement along the other (the axes line up with the ellipse).
The covariance matrix in the rotated coordinates
Now compute the covariance matrix of the rotated data Y.
| axis 1 | axis 2 | |
|---|---|---|
| axis 1 | 1.440778e+01 | 9.285502e-16 |
| axis 2 | 9.285502e-16 | 1.940396e-01 |
This covariance matrix is diagonal (up to tiny numerical rounding), which means:
- along these rotated axes, the two coordinates are uncorrelated,
- the variation in the data separates cleanly into two independent directions.
A^T=\left[\begin{array}{rrrrrrrrrrrr}2.9 & -1.5 & 0.1 & -1.0 & 2.1 & -4.0 & -2.0 & 2.2 & 0.2 & 2.0 & 1.5 & -2.5 \\ 4.0 & -0.9 & 0.0 & -1.0 & 3.0 & -5.0 & -3.5 & 2.6 & 1.0 & 3.5 & 1.0 & -4.7\end{array}\right]
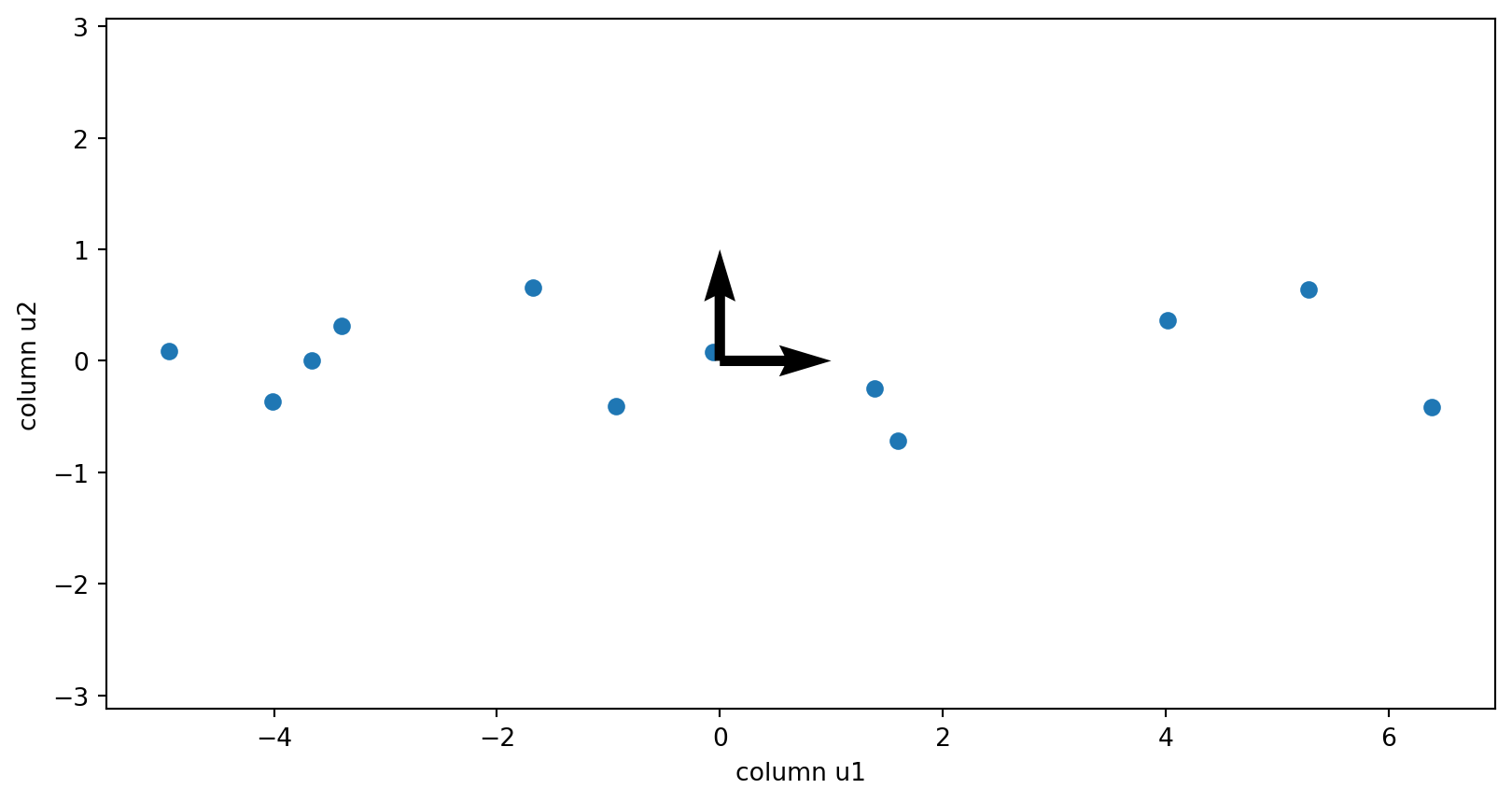
Plotting
The columns of the V matrix:
Walk through why these vectors are actually the eigenvectors of the covariance matrix.
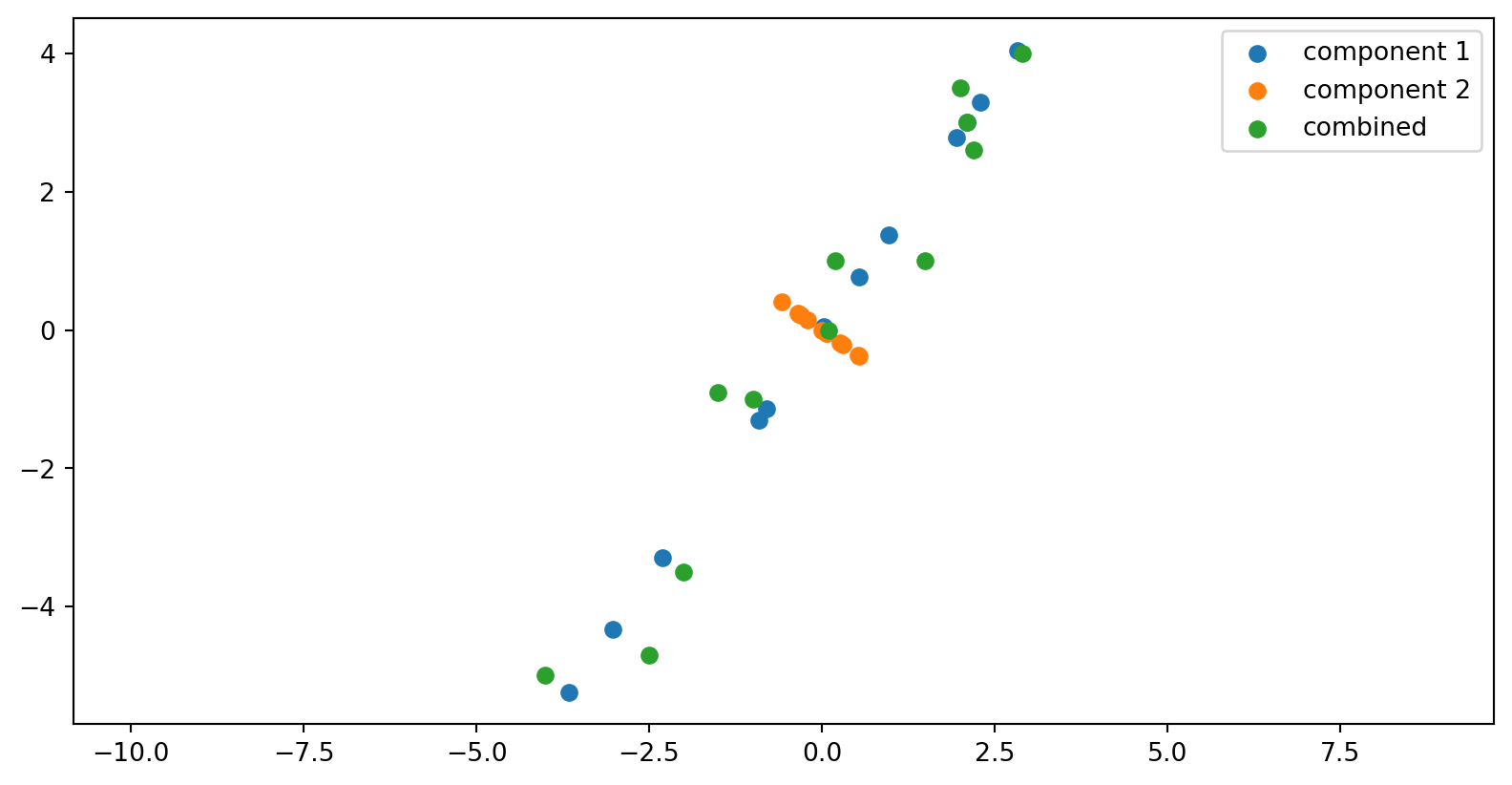
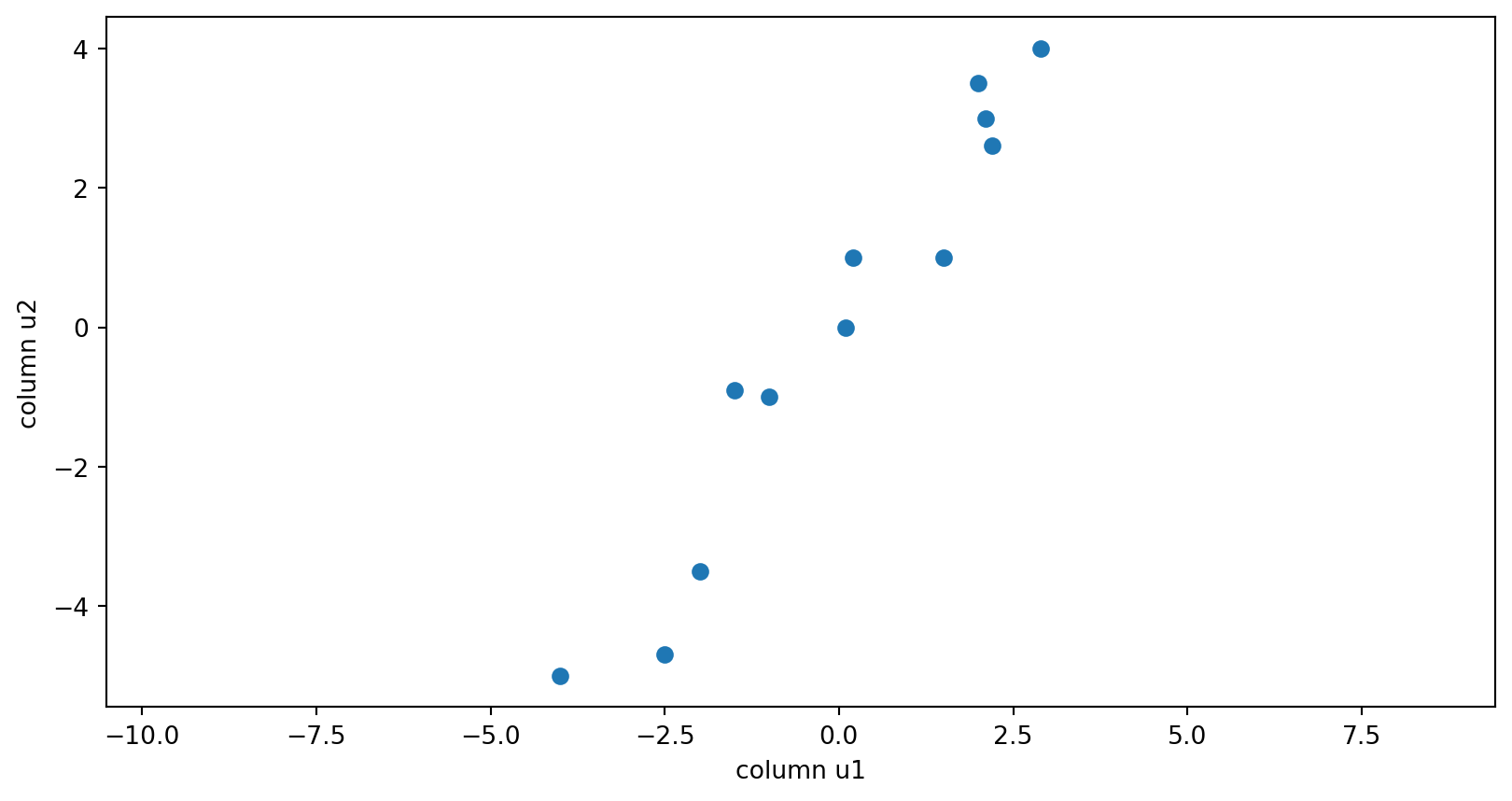
The components of the data matrix
The columns of the U matrix